Struttura genica dei procarioti e caratteristiche dei promotori batterici
Slide sulla struttura genica dei procarioti. La Pdf illustra la struttura genica dei procarioti, focalizzandosi sui promotori batterici, con diagrammi esplicativi. Questo materiale di Biologia per l'Università è utile per lo studio autonomo dei concetti chiave della genetica procariotica.
Mostra di più30 pagine
Visualizza gratis il Pdf completo
Registrati per accedere all’intero documento e trasformarlo con l’AI.
Anteprima
Struttura genica dei procarioti
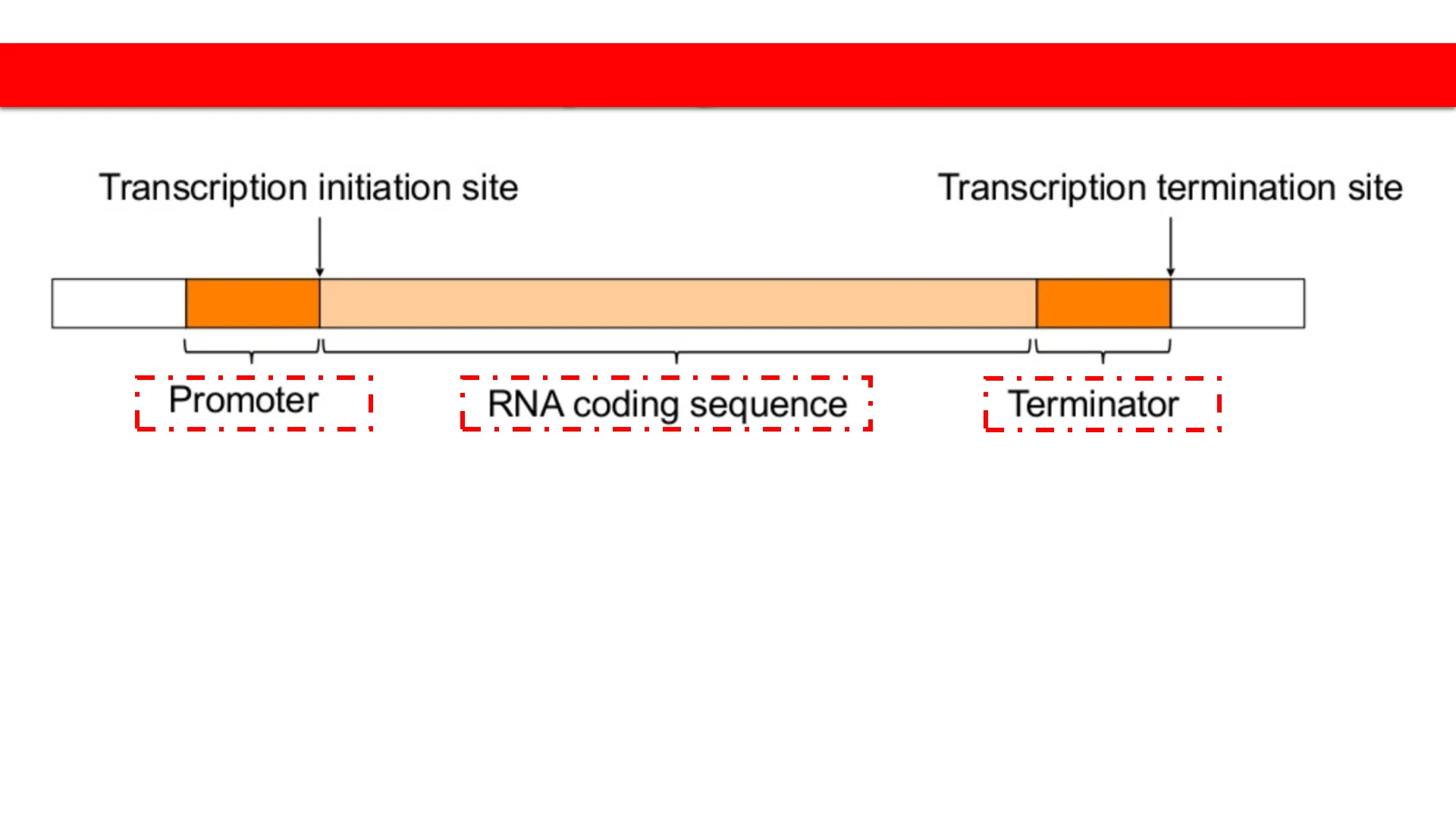
Prokaryote gene structure
Transcription initiation site
Transcription termination site
1
1
L
Promoter
I
i
RNA coding sequence
L
Terminator
I
Upstream and Downstream sequences
Caratteristiche dei promotori batterici
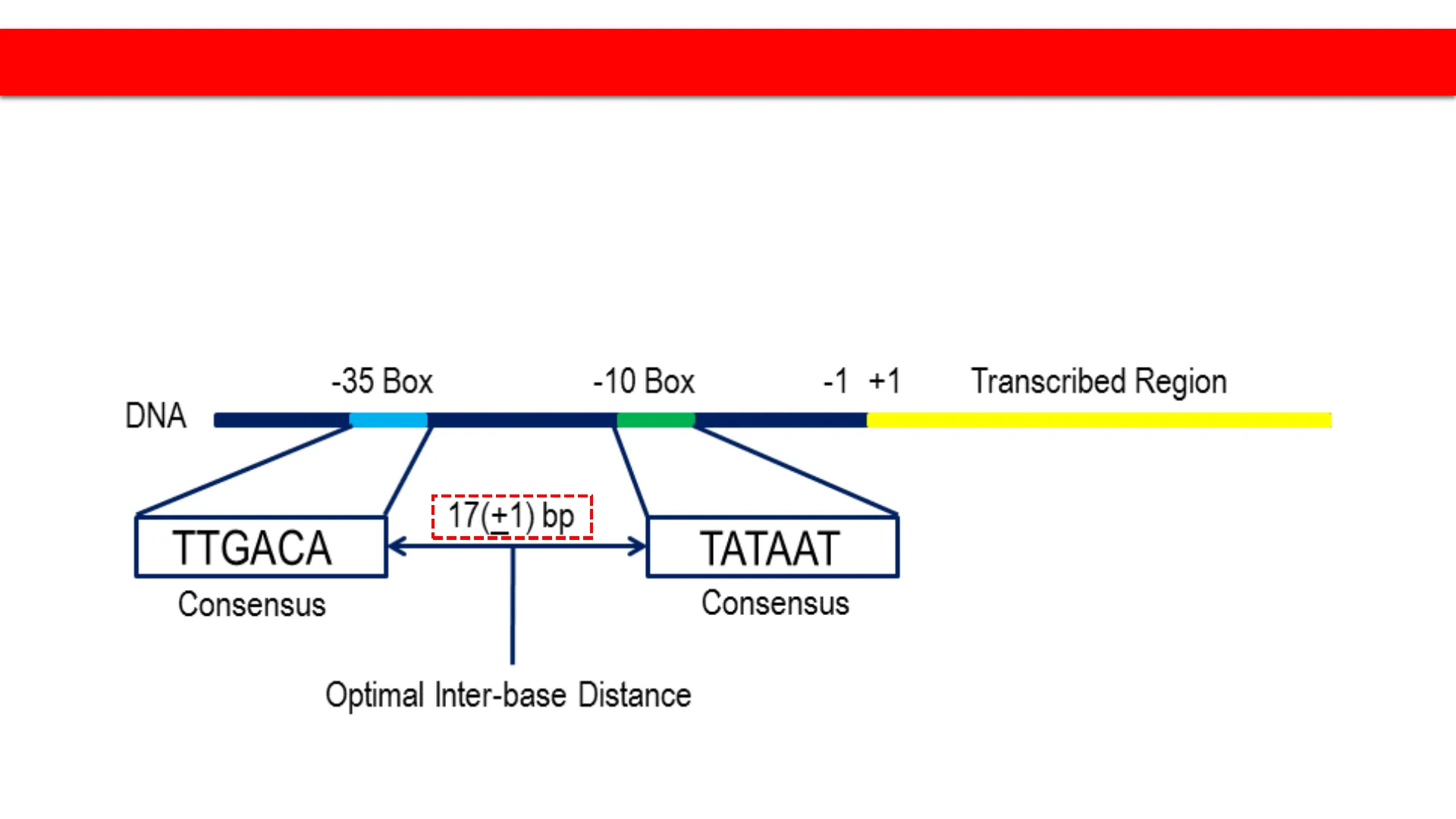
- Two conserved sequences located at -35 and -10 nt upstream
of the RNA Pol starting site - The sequences are thus called the -35 box and -10 box
-35 Box
-10 Box
-1 +1
Transcribed Region
DNA
17(+1) bp
TTGACA
TATAAT
Consensus
Consensus
Optimal Inter-base Distance
Sequenze consenso delle box -35 e -10
100
nucleotide frequency (%)
75
50-
25
0
TT GA CA
TATAAT
-35
15-19
nucleotides
-10
Elementi aggiuntivi nei promotori batterici
An additional DNA element that binds RNA polymerase is found in some
strong promoters
UP-element
-35
-10
+1
(17-19 bp)
discriminator
1
-35
-10
+1
Produzione di trascritto in base alla forza del promotore
Genes with stronger promoters produce more
transcript
gene A
gene B
DNA
TRANSCRIPTION
TRANSCRIPTION
RNA
RNA
TRANSLATION
TRANSLATION
A
AA
A
A
B
A
A
A
A
A
A
A
A
A
A
A
A
A
A
A
A
A
RNA Pol nei batteri
- Bacteria have a single RNA Pol
- In E. coli, the core enzyme has five subunits: two a, one , one B',
and one w (a2BB'w). - A o factor binds to the core enzyme forming the holoenzyme
- This factor provides the RNA Pol with promoter recognition activity
B
B'
a+a02
a2BB'
a2BB'@
(core enzyme)
Il fattore sigma media il legame della polimerasi al promotore
discriminator
"extended -10"
/
-35
-10
3
2
1
C
4.2
4.1
3.2
3.1
3.0 2.4 2.3 2.2 2.1
1.2
1.1
N
melting
Legame iniziale della RNA Pol
aNTD
aCTD
4
02
UP-element
-35
-10
The RNA Pol first binds to the 35 element and then to the -10
Transizione al complesso aperto
Transition to the open complex is mediated by
changes in the RNA Pol and in the promoter
The initial "melting" of the
DNA double-strand occurs
between positions -11 and
+2
RNA
polymerase
upstream
DNA
1
¥+1
downstream
DNA
promoter
binding
(closed
complex)
promoter
"melting"
(open
complex)
Meccanismi di transizione al complesso aperto
5'
-12
-11
-10
-9
-8
-7 -3'
T
A
T
A
A
T
-
AT ATT
A
3'
5'
no promoter melting
no transcription
This process is called
isomerization
A
T
5'
-12
10
-9< -8
-7
·3'
T
T
A
A
A
T
3'
A
T
T
A
5'
promoter melting
transcription
-11
Struttura del complesso DNA/RNA Pol
RNA polymerase
DNA exit
channel
Coding strand
3'
DNA entry
channel
5'
5'
3'
3'
rNTP entry
channel
RNA 5'
RNA exit
channel
Direction of transcription
The double helix re-
forms at -11 in the
upstream DNA
behind the enzyme
Figure 15-14
Molecular Biology: Principles and Practice
· 2012 W. H. Freeman and Company
Orientamento della RNA Pol
- The RNA Pol must transcribe the correct
DNA strand - The direction of the RNA Pol movement
determines which of the two DNA strands
is to serve as the template for
transcription - RNA Pol direction, in turn, is determined
by the orientation of the promoter
(A)
DNA double helix
1
Ctcccc cc cc CC CC CCCC
3'
5'
3'
5'
GGGGGGGGGGGGGGGGGG
5'
3'
CCCCCCC
RNA
an RNA polymerase that moves from left to right makes
RNA by using the bottom strand as a template
(B)
GGGGGGG
3'
5'
c cc cc cc cc cc cc cc ccc
5'
3'
3'
5'
GGGGGGGGGGGGGGGGGG
an RNA polymerase that moves from right to left makes
RNA by using the top strand as a template
Sequenze consenso e orientamento
-35 consensus sequence TTGACA
-10 consensus sequence TATAAT
Back
RNA Pol
Front
5'
TTGACA
AACTGT
TATAAT
ATATTA
3'
3'
5'5
3'
3'
5'
5
3'
5'
5
3'
3'
5
5'
3
5'
3
3'
Direzioni di trascrizione lungo un tratto del cromosoma batterico
Directions of transcription along a short portion of a
bacterial chromosome
RNA transcripts
DNA of E. coli chromosome
gene a
gene d
gene e
3'
3'
gene b
gene c
gene f
gene g
5'
5000 nucleotide pairs
5'
Riepilogo delle fasi di inizio della trascrizione
- The o factor binds to the promoter sequence (-10 and -35)
- Core enzyme binds to the o factor and promoter, but the DNA is
still closed (closed promoter complex) - The holoenzyme untwists the DNA double-strand (open promoter
complex) - The RNA Pol binds to the -10 sequence and to start transcribing
Requisiti per l'inizio della sintesi da parte della RNA Pol
RNA Pol does not need a primer to start synthesis
. There are still requirements for the RNA Pol to get started:
> The DNA template must be brought into the polymerase active
site and held stably
> The initiating ribonucleotide must be brought into the active
site and held stably on the template. At the same time, the next
NTP must be presented with the correct geometry for the
polymerization chemistry to occur.
. The enzyme must make specific interactions with the DNA template
strand, the initiating ribonucleotide, and the second ribonucleotide
Sintesi abortiva
Abortive Synthesis
During initial transcription, RNA Pol produces and releases short RNA
transcripts of <9 nt: abortive synthesis.
Three models have been proposed to explain the abortive synthesis:
1. Transient excursions
2. Inchworming
3. Scrunching
Modello di Scrunching
. This model proposes that DNA downstream from the stationary,
promoter-bound, polymerase is unwound and pulled into the
enzyme.
· The DNA accumulated within the enzyme is accommodated as
single-stranded bulges.
· This process generates energy
(NTP)n (PPi)n
-35
-10 +1
abortive RNA
-35 -10 +1
Fuga dal promotore
Promoter escape
. RNA Pol escapes from the promoter and enters the elongation phase only after
synthesizing a transcript of >10 nucleotides.
. A transcript >10 nucleotides cannot be accommodated within the region where it
hybridizes to the DNA and must start threading into the RNA exit channel
· The regions 3/4 of the o subunit facilitate this transition
discriminator
"extended -10"
-35
-10
4
3
2
1
C
4.2
4.1
3.2
3.1
3.0 2.4 2.3 2.2 2.1
1.2
1.1
N
melting
Rilascio del fattore sigma e transizione all'allungamento
. The o subunit eventually is released using energy accumulated
during the scrunching phase
. The o subunit release causes conformational changes in the RNA
Pol that facilitate the promoter escape and thus the transition into the
elongation phase
Release of sigma factor and
growing of RNA strand
5'
a factor
RNA
Start site
3'
5"
promoter
Elongation
5
3
Allungamento attivo
. During elongation, the RNA Pol adds
one nucleotide at a time to the growing
RNA transcript
. During this phase, the RNA Pol uses a
step mechanism and advances in a
single step a distance equivalent to a
base pair for every nucleotide it adds
to the growing RNA chain
. Therefore, the size of the bubble
remains constant
ELONGATION
RNA nucleotides
RNA
polymerase
ATCCAA
3
C
T
3' end
G
A
A
A
UC
C
A
A
A
T
A
G
G
T
T
5'
Direction of transcription
("downstream")
Template
strand of DNA
Newly made
RNA
Copyright @ Pearson Education, Inc., publishing as Benjamin Cummings.
U
T
U
CIC
5'
C
RNA Pol durante l'allungamento
Elongation
RNA polymerase
DNA
RNA
a 0 (untranslocated)
Funzioni di proofreading della RNA Pol
RNA Pol performs two proofreading functions
Pyrophosphorolytic editing
Hydrolytic editing
Terminazione
. At the end of a gene there are sequences called terminators
that trigger the RNA Pol to dissociate from the DNA and release
the RNA chain it has made.
· Bacteria have two types of termination
· Rho dependent termination
· Rho independent termination
Fattore di terminazione della trascrizione Rho
. Rho is a protein made of six identical
subunits that form its characteristic
ring-shaped structure.
. It recognizes and binds RNA as it
exits from the RNA Pol
· Specifically, Rho recognizes RNA
sequences called rut sites
Terminazione Rho-dipendente
Rho-dipendent termination
RNAP
DNA
3'
Direction of transcription
ONA
RNAP
rut
RNAP
DNA
3'
5'
RNAP
DNA
3'
5'
ATP
ADP+Pi
Terminatore Rho-indipendente
-
dyad
symmetry
DNA
3'
5'
CCCAGCCCGCCTAATGAGCGGGCTTTTTTTTGAACAAAA
GGGTCGGGCGGATTACTCGCCCGAAAAAAAACTTGTTTT
Rho-independent terminators, also called intrinsic terminators,
consist of two sequence elements:
1. a short-inverted repeat of ~20 nucleotides
2. a stretch of about eight A:T base pairs
Formazione dell'ansa a forcina nella terminazione Rho-indipendente
Rho-independent terminator
dyad
symmetry
DNA
3'
5'
CCCAGCCCGCCTAATGAGCGGGCTTTTTTTTGAACAAAA
GGGTCGGGCGGATTACTCGCCCGAAAAAAAACTTGTTTT
RNA
5'
CCCAGCCCGCCUAAUGAGCGGGCUUUUUUUU 3'
transcript folded to form
termination hairpin
A
A
U
U
G
C
A
C
G
A,U
G
C
A
C
G
U
C
G -+A,U,C
C
G -
A,U
G
C
5' CCC A
3'
G
deletion
When polymerase transcribes an inverted
repeat sequence, the resulting RNA can form
a stem-loop structure (hairpin) by base-
pairing with itself
A
-
4
Effetto dell'ansa a forcina sulla terminazione
Formation of the hairpin causes
termination by disrupting the
elongation complex
· The hairpin works as an efficient
terminator only when it is followed
by a stretch of A:U base pairs
. Under those circumstances, when
the hairpin forms, the growing RNA
chain will be held on the template at
the active site by only A:U base
pairs.
5'
3'
UUUUUUU
5'-
3' 1
UUUUUUU